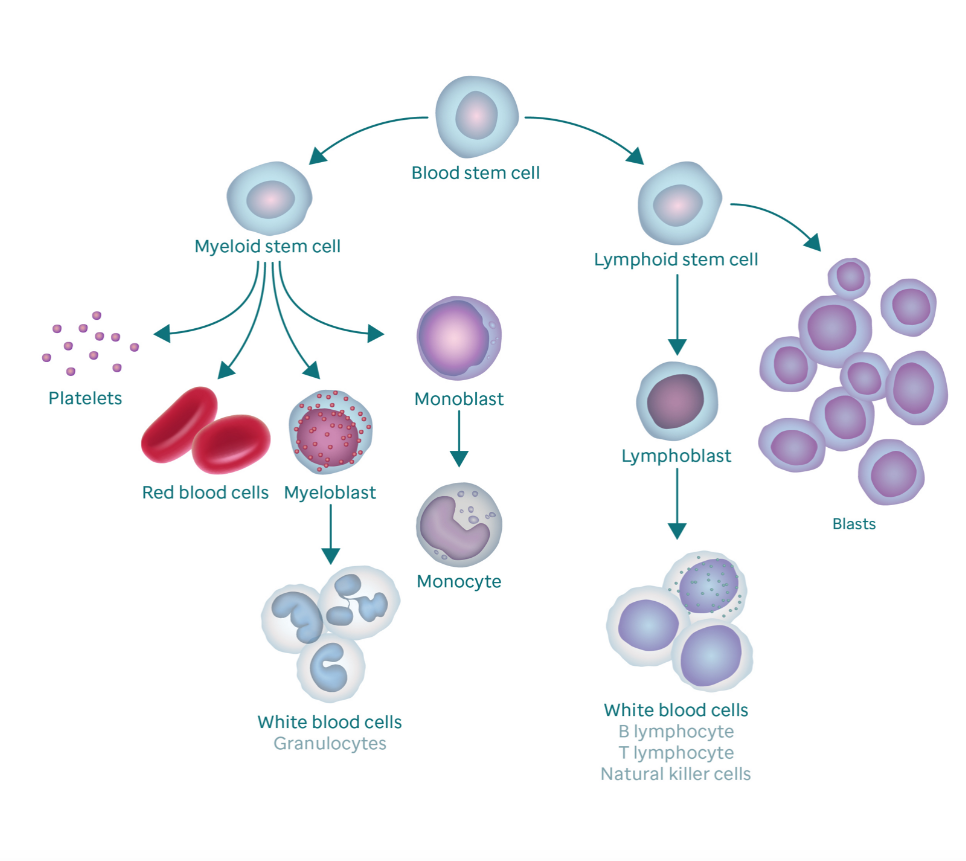
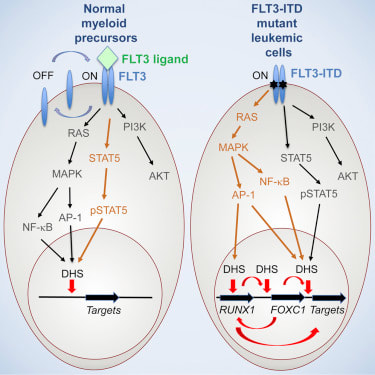
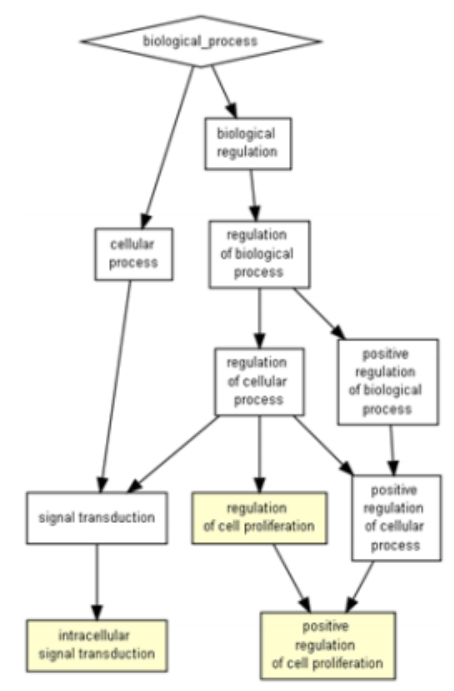
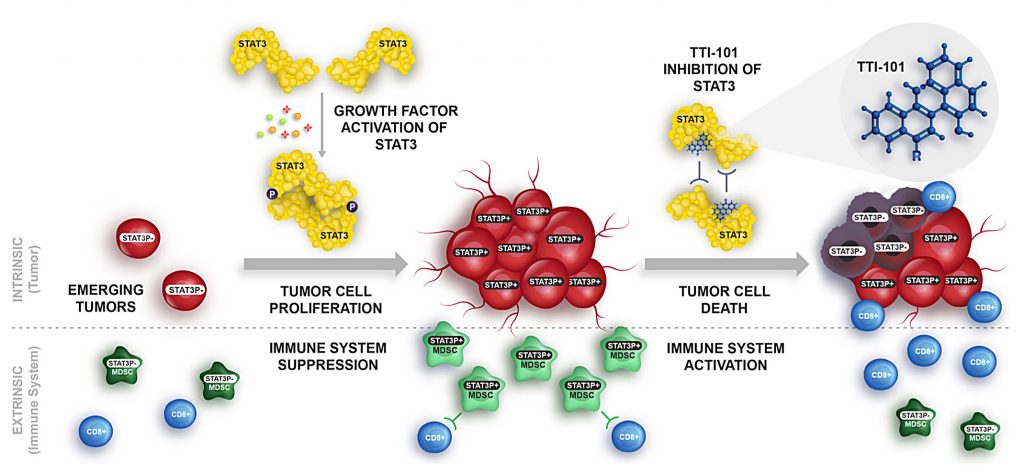
By Kylie McDanielAcute myeloid leukemia (AML) is an aggressive excess growth of immature white blood cells, or myeloblasts, within the bone marrow and blood. Due to rapid duplication of myeloblasts, the chances of metastasis, or the spread of cancer to other tissues, is significantly higher if not treated promptly (National Cancer Institute). Found in 20% of AML patients, the FLT3-ITD mutation, though rare, is closely associated with poorer prognosis (Mark Levis). FLT3 is a receptor tyrosine kinase, which is a protein responsible for cell proliferation in hematopoietic stem cells (cells from which blood cells grow), while ITD (internal tandem duplication) describes how the receptor mutated. This mutation renders the intracellular kinase domain unable to interpret surrounding stimuli as to whether the cell should grow, causing uncontrolled blood cell proliferation. The severity of this mutation lies in its role in signal transduction of hematopoietic stem cells. STAT5 proteins, a signal transducer for proliferation repression, are heavily impacted by this mutation because the ITD changes the shape of the FLT3 receptor, which prevents STAT5 proteins from both bonding to FLT3 and beginning the signal cascade, thus preventing the cell from repressing proliferation. The absence of this signal transduction is what results in an accumulation of myeloblasts that form AML (Jan-Wilhelm Kornfeld). However, a newly developed medical software called Gene Set Enrichment Analysis (GSEA) is providing oncologists with insight into the role of genetic mutations by linking groups of genes that share common biological functions, chromosomal locations, or regulations (i.e. proliferation). This is illustrated by calculating the standard deviation of every associated gene interaction the GSEA software identifies, which indicates the strength or weakness of the association between different genes. Thus, this technology is critical for doctors to choose the best inhibitor or promoter to combat the mutations present in the patient, improving their chances of remission (P. T. Aravind Subramanian). My goal was to find if there is an association between an AML patient’s cytogenetic disposition and their presence or lack of the FLT3-ITD mutation that can target a more specific and effective area for cancer therapy (which would improve prognosis). Therefore, I took 526 anonymous genetic samples from AML-positive patients in NCBI’s publicly available GEO dataset GSE14468 that were tested for the FLT3-ITD mutation and imputed them into the GSEA software to be calculated (Erdogan Taskesen). I filtered my results to focus specifically on STAT5, STAT3, and interleukin-related gene expression upregulations, or over-expressions, as these are most closely associated with instances of AML. From the 182 AML process-specific associations generated, there was an overrepresentation of positive regulation of cell proliferation, upregulation, and intracellular signal transduction of STAT5 and STAT3 proteins among the patient samples, which indicates that there is an upregulation of the gene associated with these processes. Furthermore, it was determined that inhibitors can be effectively implemented into the cancer treatment of FLT3-ITD-positive AML patients. This is an incredible breakthrough for AML patients because inhibitors directly target the mutated gene and allow for a more comfortable, less painful treatment option than chemotherapy, which is often the first line of treatment for AML patients. Additionally, these inhibitors could be improved even more to target less common pathways that are not used as often so that the effects of the treatment are not as strong as they would be otherwise due to their broadened effect on more cell pathways. This genetic identification is the key to not only the prevention of chemotherapy evasion of myeloblasts, but also to the comfort and prognosis of AML patients who demonstrate this mutation. References:
0 Comments
Leave a Reply. |
ABOUTSubmit your work to be considered for publication in the Newly Created SNHS Newspaper! CategoriesOuter Space History Biology
|